Tidy data and dealing with messy data
Objectives
Knowing about different storage formats
Knowing about the tidy data format
Be able to reformat tabular data into the tidy data format
In the previous episode we read data from nicely formatted “plain” text files. But sometimes the data is in a spreadsheet or in less nicely formatted text files. In this episode we will discuss strategies for how to work with these.
Importing data from spreadsheets
We will create a spreadsheet with the following content (only columns A and B; the actual content does not have to be exactly the same):
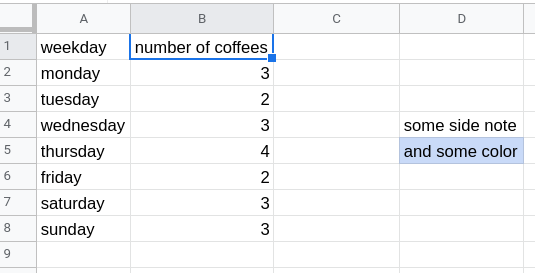
Example spreadsheet with a side note.
Copy this also to the second sheet and for demonstration purpose add some side-notes to the second sheet and also color one or two cells (some people like to give some meaning to cells using color).
Save the spreadsheet as experiment.xls.
Now we will together try to read and inspect both sheets in the Jupyter Notebook:
import pandas as pd
data = pd.read_excel('experiment.xls', sheet_name="Sheet1")
data
data = pd.read_excel('experiment.xls', sheet_name="Sheet2")
data
Discussion
We can import data from spreadsheets (more documentation)!
“Side notes” in spreadsheets can be annoying in this context.
Also encoding data in cell colors is a problem now. We will avoid those in future.
Tidy data
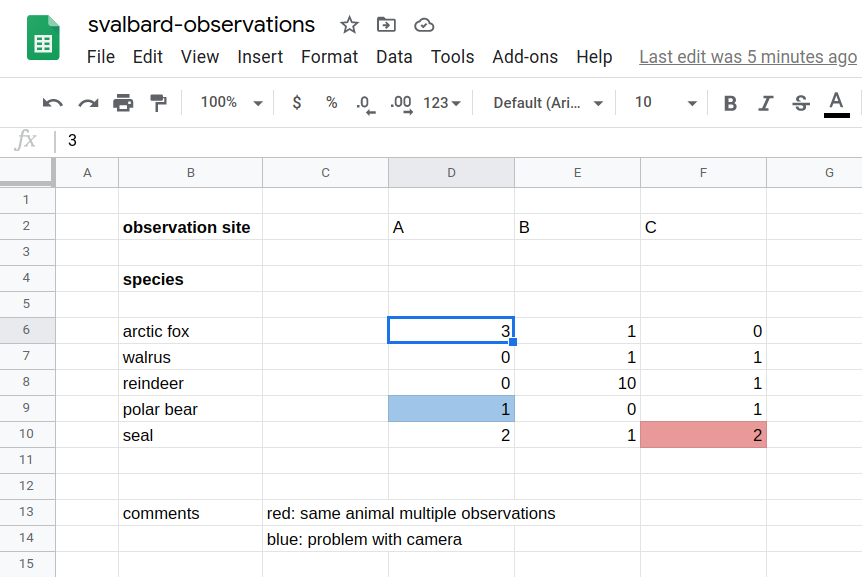
Example spreadsheet (this is a phantasy dataset, apologies to biology students/researchers - this is not my domain).
What is the problem with storing data like this?
Format: Limited interoperability with other programs
Error prone (see e.g. this famous example)
Difficult to parse (“understand”) by scripts: difficult to automate
Not in tidy format: difficult to extend/modify
How should we arrange the data?
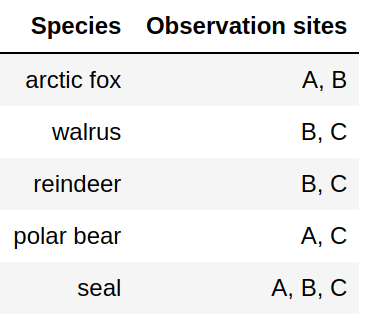
Attempt 1: Not great since we need to somehow divide at the comma. How should we deal with multiple sightings?
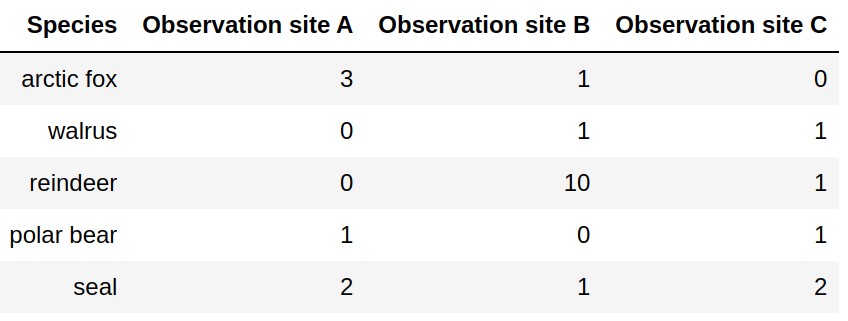
Attempt 2: Adding observation sites will force us to add columns.
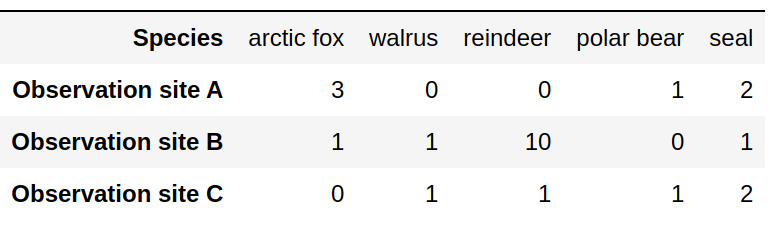
Attempt 3: Adding species will force us to add columns.
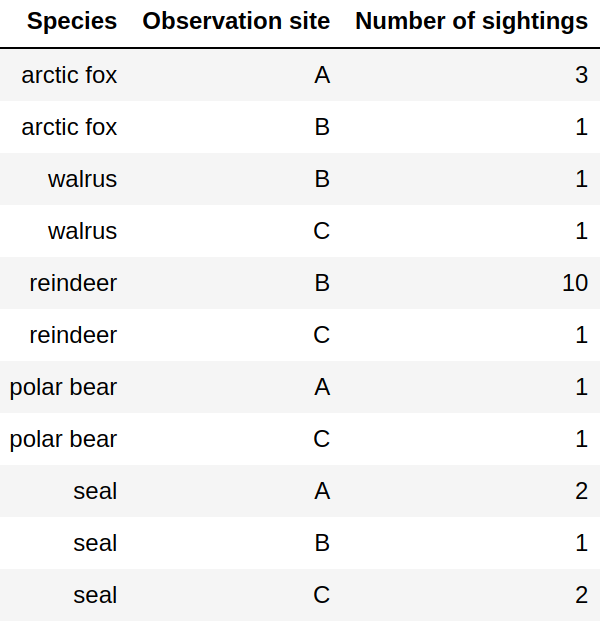
Tidy data format: Columns are variables, rows are observations/measurements. Easy to add new species and sites.
Tidy data format
Columns are variables
Rows are observations/measurements
“Long form”
Order does not matter
Easy to extend with more species and more sites without modifying the code
Structure for storing data - this does not mean that this is ideal for tables in presentations or publications
It is possible to convert between wide form and long form and back (e.g. using
pandas.meltorpandas.pivot), see this example notebook
Use a standard format
Species,Observation site,Number of sightings
arctic fox,A,3
arctic fox,B,1
walrus,B,1
walrus,C,1
reindeer,B,10
reindeer,C,1
polar bear,A,1
polar bear,C,1
seal,A,2
seal,B,1
seal,C,2
Use a format that is standard in your community, don’t invent your own
CSV is often a good choice since most visualization tools can read CSV data
There are many more formats (adapted after Python for Scientific Computing):
Name: |
Human |
Space |
Arbitrary |
Tidy |
Array |
Long term |
|---|---|---|---|---|---|---|
✅ |
❌ |
❌ |
✅ |
🟨 |
✅ |
|
❌ |
✅ |
❌ |
✅ |
❌ |
❌ |
|
❌ |
✅ |
🟨 |
✅ |
🟨 |
✅ |
|
❌ |
🟨 |
❌ |
❌ |
✅ |
❌ |
|
❌ |
✅ |
❌ |
❌ |
✅ |
✅ |
|
❌ |
✅ |
❌ |
❌ |
✅ |
✅ |
|
✅ |
❌ |
🟨 |
❌ |
❌ |
✅ |
|
✅ |
❌ |
🟨 |
❌ |
❌ |
✅ |
|
Excel |
❌ |
❌ |
❌ |
🟨 |
❌ |
🟨 |
Graph formats |
🟨 |
🟨 |
❌ |
❌ |
❌ |
✅ |
❌ |
🟨 |
❌ |
❌ |
❌ |
❌ |
Note
✅ : Good
🟨 : Ok / depends on a case
❌ : Bad
Data cleaning
Often we want to visualize data sets with inconsistent or missing entries:
Date,Organization,Number of participants
2020-09-27,UiT,20
Oct 10 2020,UiT Norges arktiske universitet,15
"Nov. 11, 2020",UiT The Arctic University of Norway,40
2020-12-12,UiT The Arctic University of Norway,-
Data cleaning is a bit outside the scope of this course (although we have done some of this in the pandas episode) but still good to know:
There are tools to clean and merge inconsistent data sets (e.g. OpenRefine, see also this Data Carpentry lesson)
This does not have to be done manually